|
||||||||||
|
||||||||||

|
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||
The vector sequence has been compiled using the information from sequence databases, published literature, and other sources, together with partial sequences obtained by Evrogen. This vector has not been completely sequenced. |
|||||||||||||||||||||||||||||||||||||||||||||||||
 Download
|
| ||||||||||||||||||||||||||||||||||||||||||||||||
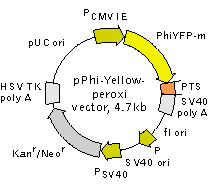
Vector description
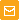
pPhi-Yellow-peroxi is a mammalian expression vector intended for yellow fluorescent labeling of peroxisomes in living cells. The vector encodes yellow fluorescent protein
Note: pPhi-Yellow-peroxi vector contains A->T substitution in position 989 (resulting in Gln to Leu substitution in amino acid position 126 of
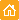
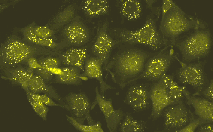 | Stably transfected T24 human bladder carcinoma cells expressing peroxisome-targeted PhiYFP-m.Image was kindly provided by Dr. Christian Petzelt (Marinpharm). |
pPhi-Yellow-peroxi vector can be used as a source of
Note: The plasmid DNA was isolated from dam+-methylated
The vector backbone contains immediate early promoter of cytomegalovirus (PCMV IE) for protein expression, SV40 origin for replication in mammalian cells expressing SV40 T-antigen, pUC origin of replication for propagation in
SV40 early promoter (PSV40) provides neomycin resistance gene (Neor) expression to select stably transfected eukaryotic cells using G418. Bacterial promoter (P) provides kanamycin resistance gene expression (Kanr) in
Expression in mammalian cells
pPhi-Yellow-peroxi vector can be transfected into mammalian cells by any known transfection method. CMV promoter provides strong, constitutive expression of the
Propagation in
Suitable host strains for propagation in
Location of features
PCMV IE: 1-589
Enhancer region: 59-465
TATA box: 554-560
Transcription start point: 583
PhiYFP-m
Kozak consensus translation initiation site: 606-616
Start codon (ATG): 613-615
A->T substitution (Gln to Leu): 989
Last amino acid in PhiYFP-m: 1312-1314
Peroxisomal targeting signal (PTS): 1315-1323
Stop codon: 1324-1326
SV40 early mRNA polyadenylation signal
Polyadenylation signals: 1534-1539 & 1563-1568
mRNA 3' ends: 1572 & 1584
f1 single-strand DNA origin: 1631-2086
Bacterial promoter for expression of Kanr gene
-35 region: 2148-2153
-10 region: 2171-2176
Transcription start point: 2183
SV40 origin of replication: 2427-2562
SV40 early promoter
Enhancer (72-bp tandem repeats): 2260-2331 & 2332-2403
21-bp repeats: 2407-2427, 2428-2448 & 2450-2470
Early promoter element: 2483-2489
Major transcription start points: 2479, 2517, 2523 & 2528
Kanamycin/neomycin resistance gene
Neomycin phosphotransferase coding sequences:
Start codon (ATG): 2611-2613
Stop codon: 3403-3405
G->A mutation to remove Pst I site: 2793
C->A (Arg to Ser) mutation to remove BssH II site: 3139
Herpes simplex virus (HSV) thymidine kinase (TK) polyadenylation signal
Polyadenylation signals: 3641-3646 & 3654-3659
pUC plasmid replication origin: 3990-4633
References:
- Gorman C. High efficiency gene transfer into mammalian cells. In DNA cloning: A Practical Approach, Vol. II. Ed. D. M. Glover. (IRL Press, Oxford, U.K.). 1985; 143-90.
- Haas J, Park EC, Seed B. Codon usage limitation in the expression of HIV-1 envelope glycoprotein. Curr Biol. 1996; 6 (3):315-24. / pmid: 8805248
- Kozak M. An analysis of 5'-noncoding sequences from 699 vertebrate messenger RNAs. Nucleic Acids Res. 1987; 15 (20):8125-48. / pmid: 3313277
Notice to Purchaser:
PhiYFP-related materials (also referred to as "Products") are intended for research use only. The Products are covered by U.S. Pat. 7,951,923; European Pat. 03779067; and other Evrogen Patents and/or Patent applications pending. By use of these Products, you accept the terms and conditions of the applicable Limited Use Label License.
|
Copyright 2002-2023 Evrogen. All rights reserved. Evrogen JSC, 16/10 Miklukho-Maklaya str., Moscow, Russia, Tel +7(495)988-4084, Fax +7(495)988-4085, e-mail:evrogen@evrogen.com |


