|
||||||||||
|
||||||||||

cDNA synthesis kit
SUPPORT |
|
|
Mint-2 cDNA synthesis kit is designed to synthesize full-length-enriched double stranded (ds) cDNA from total or poly(A)+ RNA. Synthesized cDNA can be used for construction of cDNA libraries, subtractive hybridization (SSH), high-throughput sequencing on the next generation sequencing platforms (Roche/454, ABI/SOLiD or Illumina/Solexa), and other applications. |
Technology overview
cDNA synthesis method utilizes a specific feature of the Moloney murine leukemia virus reverse transcriptase (RT). First-strand cDNA synthesis starts from the 3'-end CDS adapter containing an oligo(dT) sequence that anneals to poly(A) stretches of RNA. When RT reaches the 5'-end of the mRNA, it adds several non-template nucleotides, primarily deoxycytidines, to the 3'-end of the newly synthesized first-strand cDNA (Schmidt and Mueller, 1999; Matz et al., 1999). This oligo(dC) stretch base anneals to a complementary oligo(dG) sequence located at the 3'-end of a special deoxyribooligonucleotide adapter, called PlugOligo. Mint RT identifies PlugOligo as an extra part of the RNA template and continues first strand cDNA synthesis to the end of the oligonucleotide, thus incorporating PlugOligo sequence into the 5'-end of cDNA.
The last 3'-dG residue of the PlugOligo is a terminator nucleotide containing a 3'-phosphate group. This blocking group prevents unwanted extension of the PlugOligo. Under standard conditions Mint RT rarely uses PlugOligo as a template; however, a specially tailored IP-solution (solution for incorporation of PlugOligo sequence) dramatically increases the efficiency of this process.
At the final step, ds cDNA is amplified by PCR. Use of Encyclo polymerase and specially designed primers allows synthesis of full-length-enriched cDNA that is flanked by PlugOligo and CDS adapter sequences.
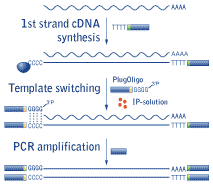 |
Schematic outline of Mint cDNA synthesis. |
|---|
cDNA prepared with Mint-2 kit can be normalized using Trimmer-2 kit (Cat# NK003) to decrease the prevalence of high abundant transcripts.
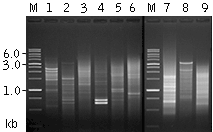 | Mint-amplified cDNA from different sources.
1 – mouse liver; 2 – mouse skeletal muscle; 3 – mouse brain; 4 – human leucocytes; 5 – human lung; 6 – human skeletal muscle; 7 – mosquito grub; 8 – copepod |
References:
- Matz M, Shagin D, Bogdanova E, Britanova O, Lukyanov S, Diatchenko L, Chenchik A. Amplification of cDNA ends based on template-switching effect and step-out PCR. Nucleic Acids Res. 1999; 27 (6):1558-60. / pmid: 10037822
- Schmidt WM, Mueller MW. CapSelect: a highly sensitive method for 5' CAP-dependent enrichment of full-length cDNA in PCR-mediated analysis of mRNAs. Nucleic Acids Res. 1999; 27 (21):e31. / pmid: 10518626
|
Copyright 2002-2023 Evrogen. All rights reserved. Evrogen JSC, 16/10 Miklukho-Maklaya str., Moscow, Russia, Tel +7(495)988-4084, Fax +7(495)988-4085, e-mail:evrogen@evrogen.com |


